Protein Network
This web page was produced as an assignment for Genetics 677, an undergraduate course at University of Wisconsin: Madison.
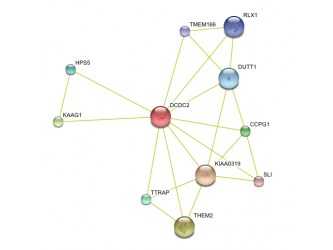
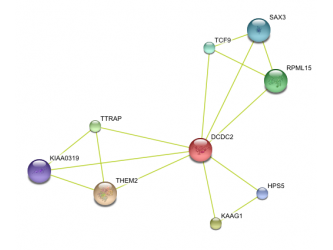
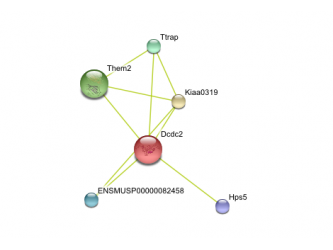
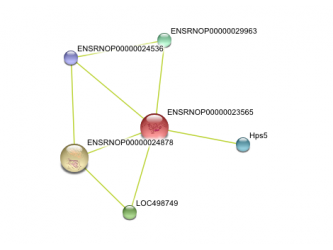
Human protein interaction network Chimp Protein interaction network Mouse protein interaction network Rat protein interaction network
Analysis
This data was retrieved from the database called String. Text mining results seem to indicate that DCDC2 gene in humans has predicted functional partners. These partners are predicted to be KIAA0319, THEM2, KAAG1, CCPG1, TTRAP, HPS5, DUTT1, RLX1, TMEM166, and SLI. The chimpanzee DCDC2 protein's functional partners are predicted to be THEM2, KAAG1,TTRAP, RPML15, TCF9, SAX3, HPS5, KIAA0319. Text mining also predicts the functional partners of mouse DCDC2. Its immediate partners are predicted to be Kiaa0319, Them2, Ttrap, ENSMUSP00000082458, and Hps5. The rat DCDC2 homolog is known as ENSRNOP00000023565. Text mining predicts its functional partners to be ENSRNOP00000024878, LOC498749, ENSRNOP00000029963, Hps5, and ENSRNOP00000024536.
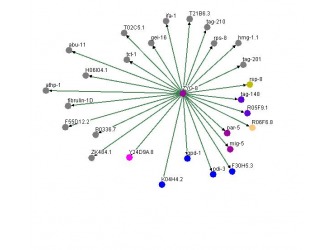
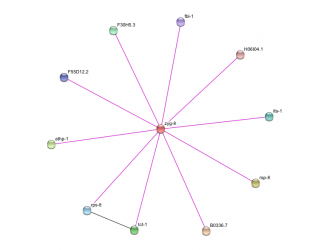
Figure 1. Osprey ZYG-8 Protein-Protein interaction Figure 2. String ZYG-8 Protein-Protein interaction
Anaylsis
The image on the right was retrieved using the database called Osprey. The image on the left was retrieved using the database called String. ZYG-8 is a zerbafish homolog of the gene DCDC2. It returned more protein-protein interactions when searching for this gene, ZYG-8, as compared to String database. However, when looking in other database within Osprey such as the mouse, rat, human, and fly databases for the DCDC2 gene and its homolog, Osprey turned up no protein-protein interactions compared to String database. This leads me to believe that when someone is researching a less known or less studied gene, a person should use both databases to find the protein-protein interactions. This is due to the fact that the two protein-protein interactions graphs are different, with one having more interactions present.