DNA Motifs
This web page was produced as an assignment for Genetics 677, an undergraduate course at University of Wisconsin: Madison.
|
Gene Data |
Figure 1. Crystal structure of the DNA binding domain of the heat shock transcription factor |
I found no motifs using full length DCDC2 DNAgene most likely because of its length.
I used DCDC2 mRNA sequence instead, and depending on the cut-off values I used, many motifs were found.
Cut-off value Number of motifs
75 275
80 218
85 (default) 149
90 81
95 25
97 15
100 7
Transfac returned 7 motifs from the 100 cut-off :
Motif Name Description Consensus
HSF heat shock factor (yeast) AGAAN
HSF heat shock factor (Drosophila) AGAAN
MATalpha2 mating factor alpha2 NCRTGTNNWN
CdxA CdxA MTTTATR
CdxA CdxA WWTWMTR
SRY sex-determining region Y gene product AAACWAM
AML-1 arunt-factor AML-1 TGTGGT
I used the mRNA sequence in MEME to find motifs in the DCDC2 gene and I identified 3 motifs.
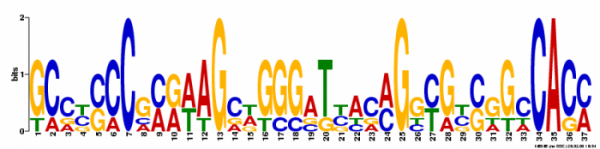
Figure 2. The first motif had an e-value of 6.7e+000 and a log likelihood ratio of 154. It had a width of 37 nucleotides and occured at 5 different sites.
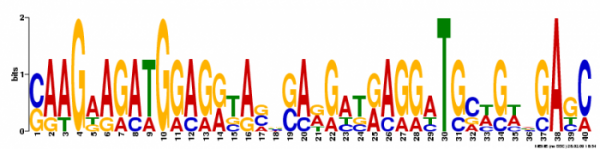
Figure 3. The second motif had an e value of 2.9e+002 and a log likelihood ratio of 154. It had a width of 40 nucleotides and occured at 5 different sites.
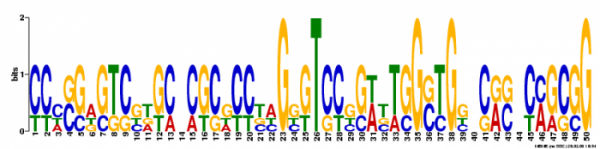
Figure 4. The third motif had an e value of 1.0e-+003 and a log likelihood ratio of 162. It had a width of 50 nucleotides and occured at 5 different sites.
Analysis
The two programs I used to determine DNA motifs were MOTIF and MEME. These two programs-MEME and MOTIF-had different formats in presenting their information and data. MOTIF and MEME allowed you to input the gene sequences, however, I put in the mRNA sequence into both since the gene sequence was too long. When I used MOTIF, it returned results that linked to a database known as Transfac, which gave a description of the type of motifs found within your sequence. I really liked having a general description that was provided with the motifs one found within the database. The one thing I did not like about MOTIF was the letters they used to represent the nucleotides within the gene.
MEME was interesting in the fact that it displayed your result in logos. I found the logos more visually pleasing then the letter representation that MOTIF used. The logos in my opinion were much easier to interpret and understand. However, MEME did not provided a list or a name of the motif(s) you were looking at, so I found it slightly more confusing and disappointing since I could not always understand what exactly I was looking at. I believe that a combination of the two programs would be your best bet in figuring out locations of the motif within the gene. That way one can further research the motifs that came up in the gene to see what proteins or transcription factors interact with your gene. In addition to it would be beneficial to double check that one is getting the same results between the two programs.
References
MOTIF
MEME figure 2,3,4
PDBSUM figure 1
Megan Holler
[email protected]
Last Updated: 2/2/09