Protein Domains
This web page was produced as an assignment for Genetics 677, an undergraduate course at University of Wisconsin: Madison.
Gene Data
|
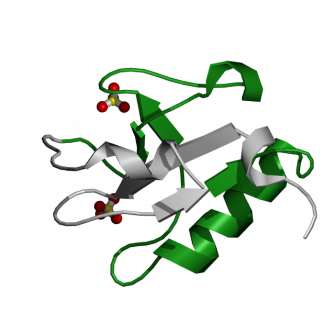
Figure 1. Structure of N-terminal doublecortin domain from DCLK: WT protein |
Pfam

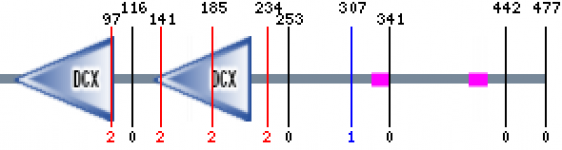
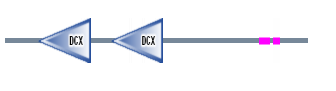
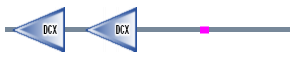
Figure 2. DCDC2 protein sequence
Pfam
When I used Pfam the DCDC2 protein sequence resulted in the protein map shown above. The domains shown above are consider to be significant in the Pfam search. The green domain represents DCDC2 N-terminus of proteins DCX which has an e-value of 4.5e-27 and 2e-17 for the first and second domains respectively. These one or two tandemly repeated copies are around the 80 amino acid region (2). The region has been suggested to bind to the first DC domain of doublecrotin binding tubulins, which enhaces microtubule polymerization(2). These two binding domains are said to be a part of the biological process: "intracellular signaling cascade". There were three other proteins brought up in the search, but each was declared as insignificant.
SMART
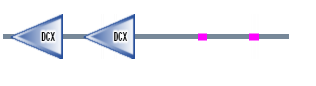
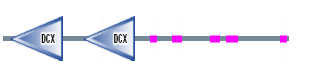
Figure 3. DCDC2 protein sequence
SMART
When I used SMART it showed the Doublecortin Gene (DCX) which are tandemly-repeated domain in doublin that are proposed to bind tubulin. Doublecortin (DCX) is mutated in human X-linked neuronal migration defects (1). The first DCX domain is in position 12 to 100 bp with an E-value of 6.28e-40 and the second DCX domain is in position 134 to 221 bp with an E-value of 3.75e-38. This can been seen in the protein above.
Prosite

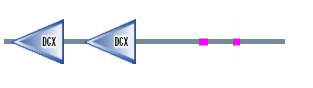
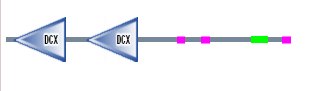
Figure 4. DCDC2 protein sequence
Prosite
Shows the same results as SMART and Pfam.
17 - 100: score = 27.542
KSVLVYRNGDPFYAGRRVVIHEKKVSSFEVFLKEVTGGVQAPFGAvRNIYTPRTgHRIRK LDQIQSGGNYVAGGQEAFKKLNYL
139 - 221: score = 23.295
CTIFLIANGDLINPASRLLIPRKTLNQWDHVLQMVTEKITLRSGAvHRLYTLEGKLVESG AELENGQFYVAVGRDKFKKLPYS
PRINTS
PRINTS did not return any results when I did a sequence search in the database.
Analysis
I found that almost all of the search engines returned that the two domains found in DCDC2 were DCX . I liked the SMART protein analysis database the best as it was easy to follow and understand. It provided a good visual image, a good written description of the DCDC2 gene, and described related protein interaction. In addition, Pfam also had good visualizations of where these domains were located and a good protein image. I liked the databases that had better visualizations rather than just presenting the written word.
The MOTIF and and PRINTS database did not return any protein domains.
Protein Domains for multiple animals
HUMAN:
My uniprot accession number gene DCDC2 for humans is Q9UHG0. In addition it gene name is DCDC2 with synonyms names as KIAA1154, and RU2 . Its protein names are listed as Doublecortin domain-containing protein 2 or Protein RU2S.
There are 2 isforms for the human DCDC2 protein produced by alternative splicing. Isform 1 has been chose as the canonical sequence Isform 2 is different from isform 1 due to missing amino acids 1-247 and amino acids 248-307 are changes from RKSKGSGNDR...KNSQETIPNS to MKMWNNWGWC...FDFHCVFVSI
The gene ontology terms for human DCDC2, found in uniprot, listed in the biological process is cellular defense response, intracellular signaling, and neuron migration.
MOUSE:
My uniport accession number for mice isQ5DU00.In addition its gene name is Dcdc2 with synonyms as Dcdc2a and Kiaa1154 . It s protein names are listed as Doublecortin domain-containing protein 2
There are 2 isforms for the mouse DCDC2 protein produced by alternative splicing. Isform 1 has been choses as the canonical sequence The isform 2 has amino acid from 308-368 change from EGIFKAGAER...ANQKEDFSAM to WLIKVERDTC...KAQPTYGHSM and has amino acids 369-475 missing.
Figure 5. Top: Left is the human protein homolog, center is the chimpanzee protein homolog, and right is the dog protein homolog.
Bottom:Left is the cow protein homolog, center is the mouse protein homolog, and right is the rat protein homolog.
Analysis
These are the protein domains of the DCDC2 gene in humans and homologs of other animals found using SMART. The two tandem DCX domains that define DCDC2 remained the same. This indicates that these two tandem repeats are most likely highly conserved through the genome within certain species throughout evolution. When using the SMART database the transmembrane segments of a protein are predicted by the program TMHMM2 and are represent by the color blue in the protein domain visual images. The coiled coil regions of a protein are determined by the program Coils2 and are represented by the color green in the protein domain visual images. The segments of low compositional complexity of a protein are determined by the program SEG and are represented by the color pink in the protein domains images. The intron positions are indicated with vertical lines showing the intron phase.
As one can see, the low complexity regions differ the most from animal to animal. Low complexity regions are regions of biased sequence composition. These regions tend to be comprised of different types of repeats and have been shown to be functionally important in some proteins. However, these regions are not well understood and are masked out to focus on globular domains within the protein domains. As one can see, there is also an area of coiled coil region located in the DCDC2 gene homolog in a mouse, which I find interesting since it is the only one.
In addition, these DCDC2 genes have different molecular wights and isoelectric points. The molecular weight and isoelectric point in the human DCDC2 protein are theoretically 52833.84 and 5.84, respectively. A chimpanzee DCDC2 protein has a molecular weight and isoelectric point of 52111.85 and 7.08, respectively . The isoelectric point is the pH value of the dispersion medium of a the protein suspension at which the protein particles do not move in an electric field.
References
Des Protes el al. Gene required for neuronal migration and involved in X-link subcortical laminar heterotopia and lissencephaly syndrome. Cell. 1998 Jan 9:92(1):51-61.1Sapir Tet al.Doublecortin mutations cluster in evolutionarily conserved functional domains.Hum Mol Genet. 2000 Mar 22;9(5):703-12.2
Pictures
Kinm MH et al. The DCX-domain tandems of doublecortin and double0like kinase, Nat Struct. Biol. 2003 May 10(5):324-33. Figure 1
Pfam figure 2
SMART figure 3
Prosite figure5
SMARTFigure 5
Megan Holler
[email protected]
Last Updated: 2/2/09