DCDC2 Protein Homologs
This web page was produced as an assignment for Genetics 677, an undergraduate course at University of Wisconsin: Madison.
|
Gene Data |
Phylogeny Tree
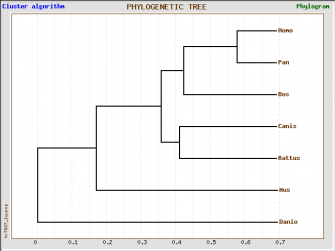
Figure 1. This tree was made out of the protein homologs using treetop. |
Pan troglodytes (chimpanzee)
Accession number: XP 527599
E-Value: 0
Canis Lupus Familiaris (dog)
Accession number: XP_853515
E-Value: 0
Bos taurus (cattle)
Accession number: XP_596512
E-Value: 0
Mus musculus (house mouse)
Accession number: NP_808245
E-Value: 5e-166
Rattus norvegicus (Norway rat)
Accession Number: XP_225316
E-Value: 0
Danio rerio (zebrafish)
Accession Number: NP_001032778
E-Value: 2e-74
Analysis
I used homologene to identify homologs of the DCDC2 protein in other organisms besides humans. I found 6 homologs. This database was very easy to use and returned results very quickly. However, this database does tend to return limited amounts of information about the genes that it finds to be similar to your gene. The lack of an E-Value in homologene is just one example where quickness precludes thoroughness. A database one can use to get an E-Value is BLAST. BLAST allows one to compare either the nucleotide or the protein sequences of the individual homolog to your gene or protein sequence. An important thing to remember when one Blasts two sequences is what type of program selection one chooses since sometimes one can get a message that says no significant similarity found. For example, on my Danio rerio sequence, I had to select the somewhat similar sequences (blastn) program in order to get an E-Value, instead of selecting the highly similar sequences (megablast) program that is normally the BLAST default setting that I used for my other five homologs.
E-values are known as expectation value. The E-value is the number of hits that would be expected to have a score equal or better than by chance alone. A good E-value would be much less than 1, and an e-value around the number 1 is what is expect by just chance alone . In principle, in order to make a decision on the significance of a match could make based on the E-value alone.
References
Homolgene http://www.ncbi.nlm.nih.gov/homologene
Picture
TreeTop Phylogenetic TreePrediction. http://www.genebee.msu.su/services/phtree_reduced.html Figure 1
Megan Holler
[email protected]
Last Updated: 2/2/09
