Conclusion and Further Directions
Results
This web page was produced as an assignment for Genetics 677, an undergraduate course at University of Wisconsin: Madison.
Results

Figure 1. Image of lettere
Inheritable dyslexia is considered a complex disease because at this point in time many genes are predicted to cause the dyslexia phenotype. An example, of the dyslexia phenotype is that children with dyslexia at a young age will have troubles distinguishing between there p,q,b,d. They will often time mix-up these letters as can seen to the right how easily this can be done (figure 1). Currently, there are 4 main genes studied, which are KIAA0319, DCDC2, Robo1, and DYX1C1; all four are involved in brain development. The gene that this website focuses on is called doublecortin domain-containing protein 2(DCDC2) which is predicted to be involved in 20% of all cases of developmental dyslexia. Developmental dyslexia often refers to biological anomalies which are usually genetic in nature that occur within the brain at various levels such as morphological or synaptic formation that occurs at the prenatal stages into childhood development. The DCDC2 gene is located on the p arm of the chromosome 6 within band 22. Through studies of the wildtype DCDC2 gene, it has been shown to modulate the migration of the neurons within the brain.
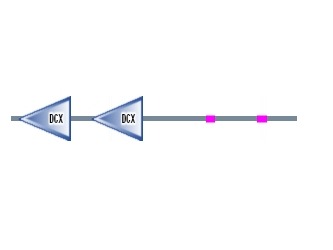
The DCDC2 gene encodes a protein that has two doublecortin peptide domains known as DCX located on the N-terminus of DCDC2. An interesting fact is that a mutated single DCX domain in humans has been known to be associated with X-linked neuronal migration defects. The DCDC2 protein has two tandem repeated copies of the DCX domain that are about 80 amino acids long(figure 2). Within this protein, these two binding domains are said to be a part of the biological process known as the intracellular signaling cascade. The first doublecortin domain of DCDC2 has been suggested to bind tubulin, whereas the role of the second doublecortin domain remains unknown. Thus, DCDC2 is involved in microtubule polymerization due to predicted binding of tubulin.
The DCDC2 gene encodes a protein that has two doublecortin peptide domains known as DCX located on the N-terminus of DCDC2. An interesting fact is that a mutated single DCX domain in humans has been known to be associated with X-linked neuronal migration defects. The DCDC2 protein has two tandem repeated copies of the DCX domain that are about 80 amino acids long(figure 2). Within this protein, these two binding domains are said to be a part of the biological process known as the intracellular signaling cascade. The first doublecortin domain of DCDC2 has been suggested to bind tubulin, whereas the role of the second doublecortin domain remains unknown. Thus, DCDC2 is involved in microtubule polymerization due to predicted binding of tubulin.
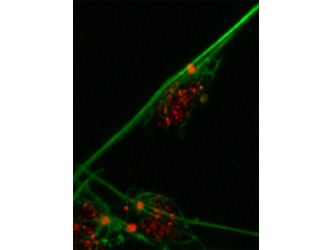
Figure 2. DCDC2 human protein domains Figure 3. Migration of neurons
The WT DCDC2 controls migration of the neurons within the reading center of the brain where it is normally expressed(figure 3). A deletion occurs within the DCDC2 gene that is associated with this. It has been found to be a large polymorphic deletion within the intron site 1of DCDC2 that is responsible for the transcription factor binding site. In addition to a deletion within the intron site 2 of DCDC2 which is also responsible for the transcription factor binding site. It has been shown that 1 out 5 dyslexics have the mutated form of the DCDC2 protein. This is due to alternative splicing that happens within the gene. Thus, DCDC2 has two isoforms. The majority of the research is focused on the intron 2 deletion. This mutation creates a less active form of the DCDC2 protein. This alteration that occurs with DCDC2 has been shown to disrupt the development of the brain circuits that are necessary for fluent reading. This can later affect the radial neuronal migration within the developed brain since DCDC2 is also down regulated. In order to take a closer look at the DCDC2 protein, especially localization, scientists have studied homologs of DCDC2 in mice.
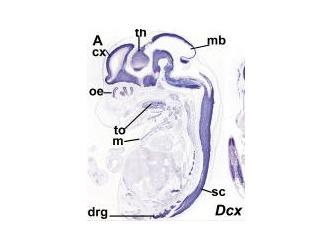
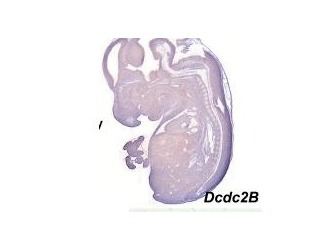
Figure 4. Shows expression profile of just the a DCX domain and DCDC2 mouse homolg DCDC2B
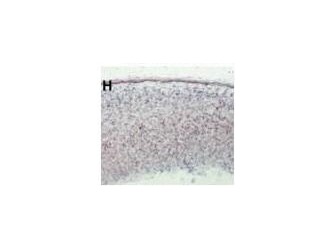
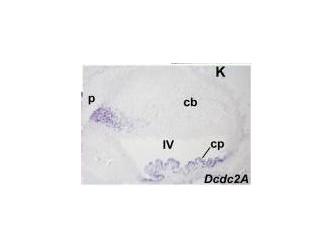
There is a paper that analyzed different expression of genes that had the DCX domain including DCDC2 within the mouse model. The blue stain in these images denotes expression of the gene whose name is indicated in each panel(3). These two images are sections of whole embryos of DCX and DCDC2B(3). The individual DCX domain is up so you can see the difference between the expression profile "snapshot" between DCDC2 and the paired DCX domains and the individual DCX domain (Figure 4)(3). As you can see, the DCDC2 image shows ubiquitous expression of DCDC2B while the DCX shows expression that is more specific and localized(3). Then, the authors decided to analyze expression in the developing neocortex which can be seen in figure 5(3). It shows expression of DCDC2B is more pronounced in its ventricular zone of the neocortex(3). Next, the authors investigated DCDC2A expression in the CNS, as seen in figure 6, which shows DCDC2A is restricted to a group of scattered neurons in the lateral-most part of the developing cerebellum(3). DCDC2A is also expressed in the choroid plexi. The authors concluded and agreed with the conclusion that DCDC2 is expressed within the entorhinal cortex, inferior temporal cortex, medial temporal cortex, hypothalamus, amygdala, and hippocampus where fluent reading processes occur because DCDC2 controls the the migration of the necessary neurons. Besides just studying the expression of DCDC2 in the brain because it is associated with neurons, researchers have also studied the DCDC2 homolog within C.elegans.
Figure 5. DCDC2B expression in developing neocortex Figure 6. DCDC2a expression in CNS
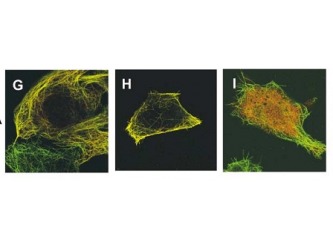
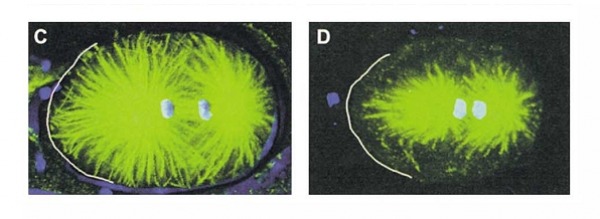
Figure 7. WT C.elegan embryo. Figure 8. Zyg-8 mutant embryo. Figure 9. Shows localization of Zyg-8 where G is both
Zyg-8 domains, H is just DCX domain alone,
I is just the kinase domain alone.
Zyg-8 domains, H is just DCX domain alone,
I is just the kinase domain alone.
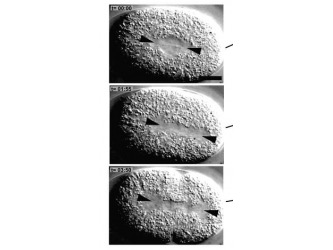
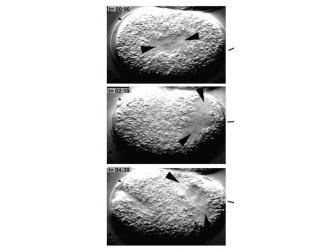
The C.elegans homolog is known as Zyg- 8. The Zyg-8 gene encodes a protein with a doublecortin-like domain and a protein kinase domain(1). It is presented at low levels in the cytoplasm throughout the cell cycle and detectable at centrosomes during prophase(1). Zyg-8 is also known to concentrate in areas of high microtubule density like the spindle and spindle poles during mitosis(1). A mutation within Zyg-8 impairs mitosis at the one cell embryonic stage(1). In the wild-type C.elegans embryos, the spindle positioning follows from two processes. First, the two centrosomes and associated pro-nuclei move to the cell center while undergoing a 90° rotation(figure 7 top)(1). As a result, the spindle sets up in the cell center and along the AP axis. Second, during anaphase, the posterior spindle pole is displaced slightly toward the posterior(figure 7 middle)(1). As a result, the spindle is asymmetrically positioned along the AP axis, leading to unequal cleavage into a larger anterior blastomere and a smaller posterior one(figure 7 bottom)(1). However, the Zyg-8 mutant fails to show proper asymmetry in embryos(1). Instead a vast majority of these mutant embryos have a spindle that was displaced in an exaggerated manner closer towards the posterior end than during normal anaphase (figure 8)(1).
Thus, when the microtubules were analyzed during anaphase the astral microtubules were shorter in the mutant as compared to the wild type Zyg-8 gene (figure 10)(1). Thus, wildtype Zyg-8 function promotes microtubule assembly during anaphase in order to ensure proper spindle positioning and asymmetric cell division(1). The authors then identified the molecular nature of Zyg-8 and they found that it encodes a doublecortin domain and a kinase domain(1). Zyg-8 was shown to be a microtubule associated protein (MAP). The authors then figured out which of the two domains caused localization to the to the microtubules(1). The kinase domain from the Zyg-8 gene alone localizes to the cytoplasm (figure 9I) and the DCX domain alone localizes to the microtubules( figure 9H)(1). This Zyg-8 gene leads to protection against depolymerization of microtubules.
Thus, when the microtubules were analyzed during anaphase the astral microtubules were shorter in the mutant as compared to the wild type Zyg-8 gene (figure 10)(1). Thus, wildtype Zyg-8 function promotes microtubule assembly during anaphase in order to ensure proper spindle positioning and asymmetric cell division(1). The authors then identified the molecular nature of Zyg-8 and they found that it encodes a doublecortin domain and a kinase domain(1). Zyg-8 was shown to be a microtubule associated protein (MAP). The authors then figured out which of the two domains caused localization to the to the microtubules(1). The kinase domain from the Zyg-8 gene alone localizes to the cytoplasm (figure 9I) and the DCX domain alone localizes to the microtubules( figure 9H)(1). This Zyg-8 gene leads to protection against depolymerization of microtubules.
There are no medications or drugs so far discovered that can treat dyslexia since it so complex and much about it remains unknown. In addition there is no known chemical in chembank that is associated with the DCDC2 gene or protein. In the future, I would take a look at the DCX domain within other genes that have this domain to see if one could find a known chemical that many disrupt microtubule or neuron formation. The complexity of these studies is why the only treatment so far is remedial education. However, dyslexia is not as simple as one gene, one disease. There are many protein-protein interactions and other genes known to be involved in dyslexia as well. Some of the most interesting protein-protein interactions found so far are more focused on the general expression of DCDC2 effects on the brain. I found these three proteins are key: CCPG1, DUTT, and SHC2(7). CCPG1, which is known as the dyslexia susceptibility 1 candidate gene 1 protein, is a chromosomal aberration that occurs involves the protein DYX1C1 that is known as another cause of dyslexia(7). DUTT is a roundabout homolog 1 precursor to the gene known as robo-1(7). DUTT is a receptor for the protein ligands SLIT1 and SLIT2, which are thought to act as molecular guidance cues in cellular migration that includes axonal navigation at the ventral midline of the neural tube and projection of axons to different regions during neuronal development(7). In the axon growth cones, the silencing of the attractive effect of NTN1 by SLIT2 may be require the formation of a ROBO1-DCC complex(7). SHC2 is another protein which is a signaling adapter that couples activated growth factor receptors to signaling pathways in neurons(7). It is involved in the signal transduction pathways of neurotrophin-activated Trk receptors in cortical neuron(7).
| the_evolving_doublecortin_dcx_superfamily.pdf | |
| File Size: | 3117 kb |
| File Type: | |
| zyg-8_a_gene_required_for_spindle_positioning_in_c.elegans.pdf | |
| File Size: | 968 kb |
| File Type: | |
References:
Gonczy, P et al (2001).Zyg-8, a Gene Required for Spindle Positioning in C. elegans, Encodes a Doublecortin-Related Kinase that Promotes Microtubule Assembly. Developmental Cell. 1,363–375. 1
Johannes, Marc (February 1 2001). Functional Brain Mapping.. May 15 2009 from http://www.lems.brown.edu/~msj/summary /summary.html. 2
Meng, H et al. (2005) DCDC2 is associated with the reading disability and modulates neuronal development in the brain. PNAS. 102(47), 17053-17058. 3
Reiner, O et al. (2006). The evolving doublecortin (DCX) superfamily. BMC Genomics. 7(188) Retrieved from http://www.biomedcentral.com/1471-2164/7/188. 4
Roth, Mark (2007). Dyslexia begins when the wires don't meet. Post-Gazette. Retrieved May 15 2009 from http://www.post-gazette.com/pg/07042/760823-114.stm. 5
The International Dyslexia Association. (2008). Dyslexia Basics. IDA. Retrieved May 15, 2008 from http://www.interdys.org/ewebeditpro5/upload/Basics_Fact_Sheet_5-08-08.pdf 6
STRING 7
Images:
(Figure 1) http://www.hopkinsvilledyslexia.org/news.html
(Figure 2) Retrieved from SMART
(Figure 3) Didkovsky, N. Hatten Lab Project
(Figure 4) Reiner, O et al. (2006). The evolving doublecortin (DCX) superfamily. BMC Genomics. 7(188) Retrieved from http://www.biomedcentral.com/1471-2164/7/188.
(Figure 5) Reiner, O et al. (2006). The evolving doublecortin (DCX) superfamily. BMC Genomics. 7(188) Retrieved from http://www.biomedcentral.com/1471-2164/7/188.
(Figure 6) Reiner, O et al. (2006). The evolving doublecortin (DCX) superfamily. BMC Genomics. 7(188) Retrieved from http://www.biomedcentral.com/1471-2164/7/188.
(Figure 7) Gonczy, P et al (2001).Zyg-8, a Gene Required for Spindle Positioning in C. elegans, Encodes a Doublecortin-Related Kinase that Promotes Microtubule Assembly. Developmental Cell. 1,363–375
(Figure 8) Gonczy, P et al (2001).Zyg-8, a Gene Required for Spindle Positioning in C. elegans, Encodes a Doublecortin-Related Kinase that Promotes Microtubule Assembly. Developmental Cell. 1,363–375
(Figure 9) Gonczy, P et al (2001).Zyg-8, a Gene Required for Spindle Positioning in C. elegans, Encodes a Doublecortin-Related Kinase that Promotes Microtubule Assembly. Developmental Cell. 1,363–375
(Figure 10) Gonczy, P et al (2001).Zyg-8, a Gene Required for Spindle Positioning in C. elegans, Encodes a Doublecortin-Related Kinase that Promotes Microtubule Assembly. Developmental Cell. 1,363–375
Gonczy, P et al (2001).Zyg-8, a Gene Required for Spindle Positioning in C. elegans, Encodes a Doublecortin-Related Kinase that Promotes Microtubule Assembly. Developmental Cell. 1,363–375. 1
Johannes, Marc (February 1 2001). Functional Brain Mapping.. May 15 2009 from http://www.lems.brown.edu/~msj/summary /summary.html. 2
Meng, H et al. (2005) DCDC2 is associated with the reading disability and modulates neuronal development in the brain. PNAS. 102(47), 17053-17058. 3
Reiner, O et al. (2006). The evolving doublecortin (DCX) superfamily. BMC Genomics. 7(188) Retrieved from http://www.biomedcentral.com/1471-2164/7/188. 4
Roth, Mark (2007). Dyslexia begins when the wires don't meet. Post-Gazette. Retrieved May 15 2009 from http://www.post-gazette.com/pg/07042/760823-114.stm. 5
The International Dyslexia Association. (2008). Dyslexia Basics. IDA. Retrieved May 15, 2008 from http://www.interdys.org/ewebeditpro5/upload/Basics_Fact_Sheet_5-08-08.pdf 6
STRING 7
Images:
(Figure 1) http://www.hopkinsvilledyslexia.org/news.html
(Figure 2) Retrieved from SMART
(Figure 3) Didkovsky, N. Hatten Lab Project
(Figure 4) Reiner, O et al. (2006). The evolving doublecortin (DCX) superfamily. BMC Genomics. 7(188) Retrieved from http://www.biomedcentral.com/1471-2164/7/188.
(Figure 5) Reiner, O et al. (2006). The evolving doublecortin (DCX) superfamily. BMC Genomics. 7(188) Retrieved from http://www.biomedcentral.com/1471-2164/7/188.
(Figure 6) Reiner, O et al. (2006). The evolving doublecortin (DCX) superfamily. BMC Genomics. 7(188) Retrieved from http://www.biomedcentral.com/1471-2164/7/188.
(Figure 7) Gonczy, P et al (2001).Zyg-8, a Gene Required for Spindle Positioning in C. elegans, Encodes a Doublecortin-Related Kinase that Promotes Microtubule Assembly. Developmental Cell. 1,363–375
(Figure 8) Gonczy, P et al (2001).Zyg-8, a Gene Required for Spindle Positioning in C. elegans, Encodes a Doublecortin-Related Kinase that Promotes Microtubule Assembly. Developmental Cell. 1,363–375
(Figure 9) Gonczy, P et al (2001).Zyg-8, a Gene Required for Spindle Positioning in C. elegans, Encodes a Doublecortin-Related Kinase that Promotes Microtubule Assembly. Developmental Cell. 1,363–375
(Figure 10) Gonczy, P et al (2001).Zyg-8, a Gene Required for Spindle Positioning in C. elegans, Encodes a Doublecortin-Related Kinase that Promotes Microtubule Assembly. Developmental Cell. 1,363–375