Phenotypes
This web page was produced as an assignment for Genetics 677, an undergraduate course at University of Wisconsin: Madison.
Mutations in the DCDC2 gene causes a reading disorder in humans that is associated with dyslexia. A question that is often asked is what phenotypes would be found in other organisms besides humans. This question can be explored with either the technique of RNA interference (RNAi) or through the use of Small Molecules to inhibit that protein or gene function. The model organism I looked at was C.elegans, with RNA interference (RNAi) used to knock down gene expression
Wormbase
In order to find a phenotype in C.elegans of my gene, I did a blast/blat search of the DCDC2 gene in Wormbase. The parameters I used were my protein sequence with the query being protein blast with an e-value threshold of 1E+0. This returned 6 hits: Y79H2A.11, F27C1.13, W07G1.1, C06C3.3, W07G1.5b, W07G1.5a. Out of the six hits only Y79H2A.11, W07G1.5b, W07G1.5a proteins had a doublecortin domain. The other three did not have descriptions of the domain present and were not further researched; however, the other three did have the GO term intracellular signaling cascade (IEA) which is one of the GO terms associated with DCDC2. These two genes that have a doublecortin domain like gene DCDC2 are Zyg-8 andW07G1.5. The Zyg-8 gene encodes a protein with a doublecortin-like domain and a protein kinase domain while mutations of Zyg-8 impair mitosis at the one cell embryonic stage (1). At the one cell embryonic stage it is normally asymmetric in embryos; however, the Zyg-8 mutant fails to show the proper asymmetry in embryos (1). The Zyg-8 is known to concentrates in areas of high microtubule density like the spindle and spindle poles during mitosis (1). It is also present at low levels in the cytoplasm throughout the cell cycle and detectable at centrosomes during prophase (1).
TheW07G1.5 gene is a microtubule assembly protein doublecortin that contains DCX domain.
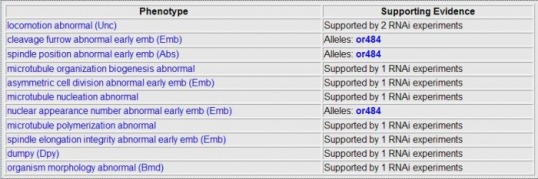
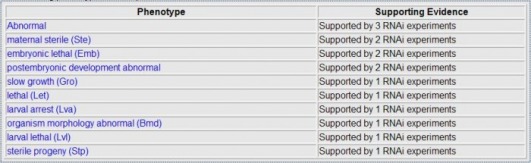
Wormbase allows one to look the phenotypes seen when RNAi is used. One can see the phenotypes for genes ZyG-8 and W07G1.5 below. W07G1. 5 did not have any phenotypes seen in any RNAi experiments
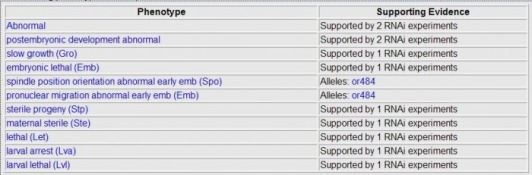
In addition, Wormbase also lists phenotypes not observed in an RNAi experiment. One can see the phenotypes not seen for genes ZYG-8 and W07G1.5.
PubChem, ChemBank, and DrugBank
Small Molecules can be used to either inhibit or activate a function of the protein known to be part of the pathway you are examining.
I would have liked to find Small Molecules that either inhibit or activate the DCDC2 gene products, but there are currently no drugs or known molecules to be involved in with DCDC2 gene product due to its location and expression in the brain. Thus, I was not able to find hits in the databases PubChem, ChemBank, or DrugBank when searching for Small Molecules involved with DCDC2 or dyslexia.
RNAi Database
RNAi database is very useful for finding movies of your phenotype. I wanted to see if I could find any movies involving the genes Zyg-8 and W07G1.5. I did find some for gene Zyg-8 and none for W07G1.5. Please see the movies below.
MOVIES:
Wildtype Embryo phenotype. Zyg-8 Mutant Embryo phenotype: 1 2 3 4 5 6 7 8
Analysis:
The E-value and score bits for gene Zyg-8 was 3e-13 and 73, respectively while gene W07G1.5 was .015 and 38 respectively. This means that gene Zyg-8 is currently the better homolog out of the two genes that is more closely related to the human DCDC2 gene. When comparing the Wormbase data to the RNAi database, I found Wormbase to be more user-friendly. It was laid out much better than the RNAi database and its links were much easier to follow. The phenotypes were much easier to find and understand what you were looking at. However, I found that RNAi database had movies of the phenotypes even though the phenotypes were not labeled clearly, and it was easier to find primary literature sources for you information and what experiments were preformed. For example, the gene W07G1.5 did not list where any of the information and evidence came from while the RNAi database did list where the experiments and information was from. This was the biggest difference I saw between the two databases. I think that both have their perks and failures, but throughmy work with my gene DCDC2 I have discovered that they work really well together.
References:
Gonczy P, Bellanger JM, Kirkham M, Pozniakowski A, Baumer K, Phillips JB, Hyman AA.
2001. zyg-8, a gene required for spindle positioning in C. elegans, encodes a Doublecortin-related kinase that promotes microtubule assembly. Developmental Cell 1:363-375.1
Figures:
(Figure 1) http://www.wormbase.org/db/gene/gene?name=WBGene00006993;class=Gene
(Figure2) http://www.wormbase.org/db/gene/gene?name=WBGene00012332;class=Gene
(Figure3) http://www.wormbase.org/db/gene/gene?name=WBGene00006993;class=Gene
(Figure4) http://www.wormbase.org/db/gene/gene?name=WBGene00012332;class=Gene
Movies:
C.elegan wildtype movie retrieved from http://www.wormclassroom.org/Modules/CellLineage/CACE/CEE.html on 5/13/09
C. elegan Zyg-8 mutant movies retrieved in 5/13/09 from http://worm-srv2.mpi-cbg.de/ceimages/331/331-E12-w16016.0.mov 1
http://worm-srv2.mpi-cbg.de/ceimages/331/331-E12-w38057.0.mov 2
http://worm-srv2.mpi-cbg.de/ceimages/331/331-E12-w57252.0.mov 3
http://worm-srv2.mpi-cbg.de/ceimages/331/331-E12-w6.mov 4
http://worm-srv2.mpi-cbg.de/ceimages/331/331-e12-1-w1.mov 5
http://worm-srv2.mpi-cbg.de/ceimages/331/331-e12-1-w2.mov 6
http://worm-srv2.mpi-cbg.de/ceimages/331/331-e12-1-w3.mov 7
http://worm-srv2.mpi-cbg.de/ceimages/331/331-e12-1-w5.mov 8
Megan Holler
[email protected]
Last Updated: 2/12/09